Active Research Projects
The Control of Transcription Factor Specificity and Activity
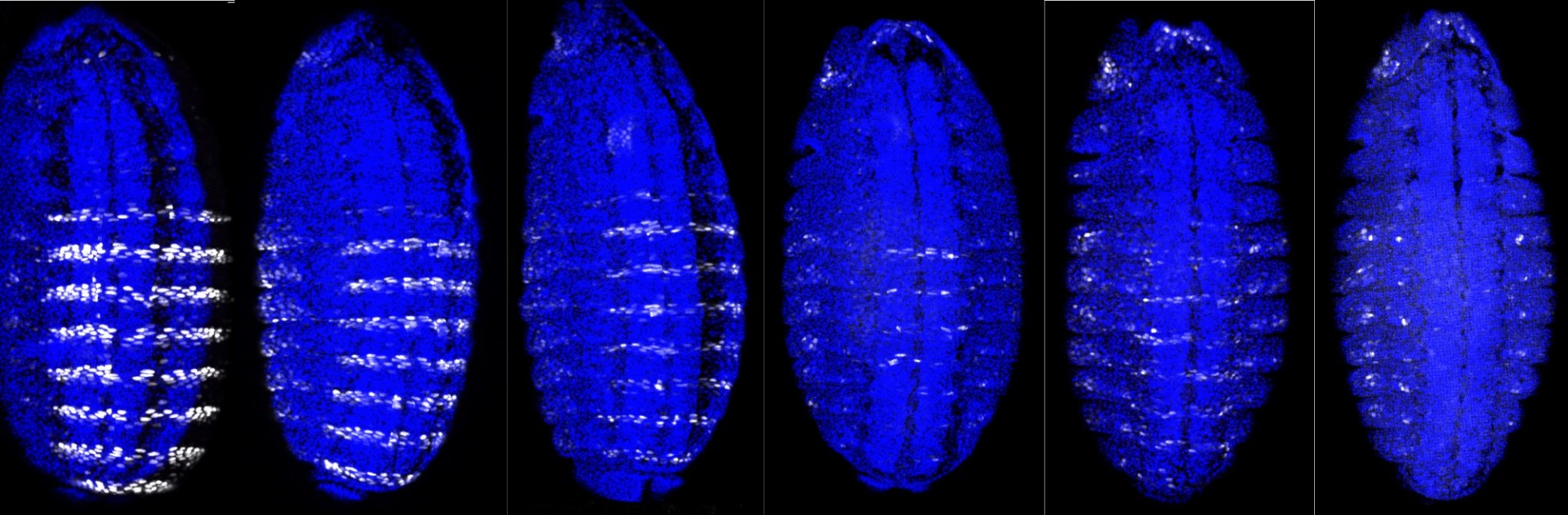
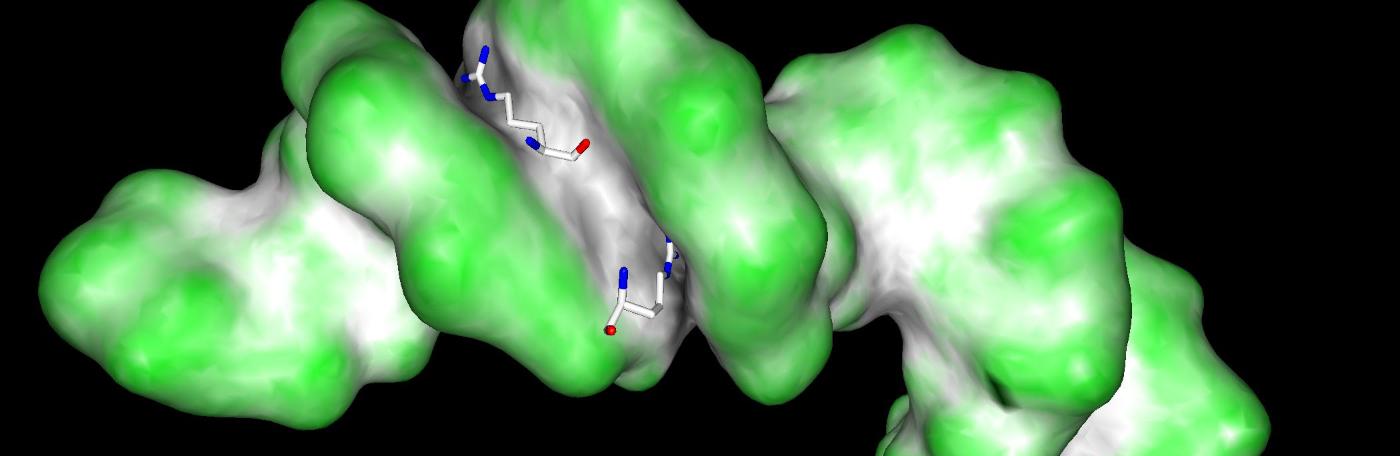
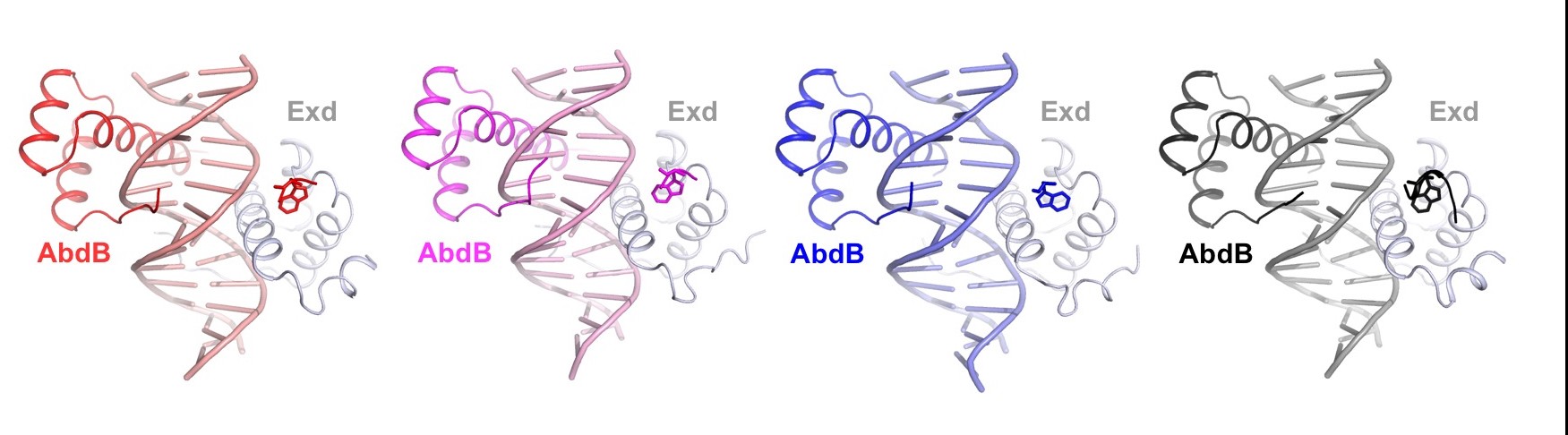
We study how transcription factors bind to the correct DNA sequences and regulate the correct target genes in vivo, with a focus on the Hox family of homeodomain proteins.
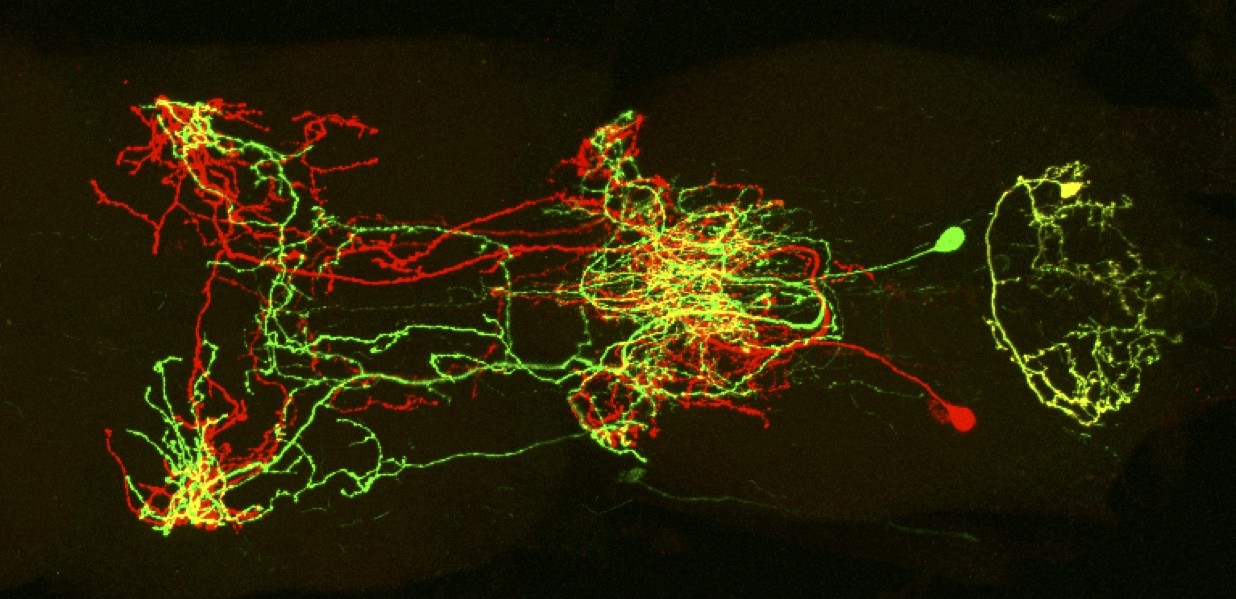
Motor Neuron Development and Circuitry
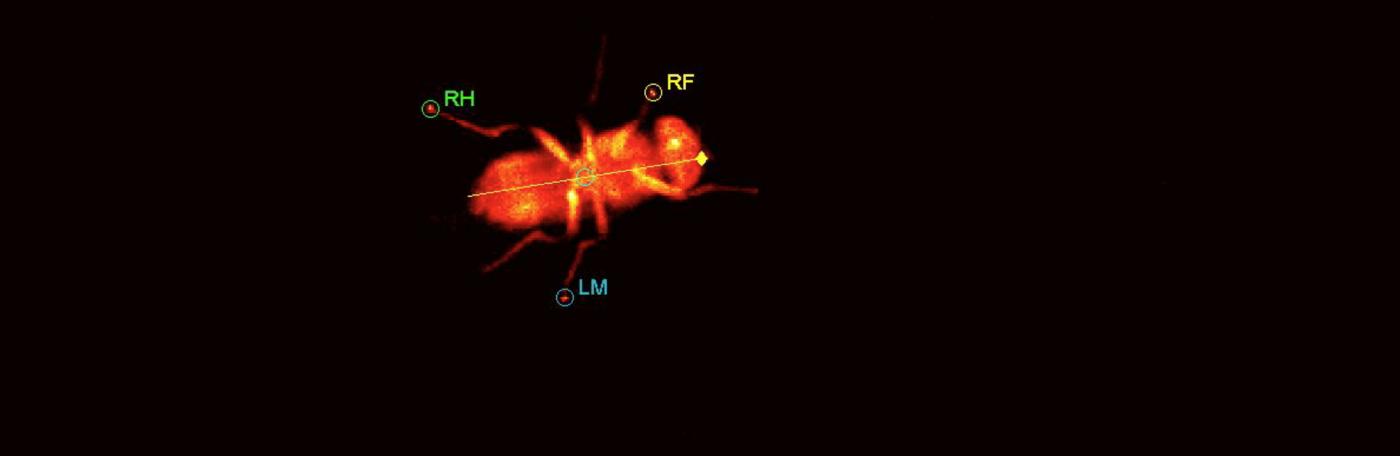
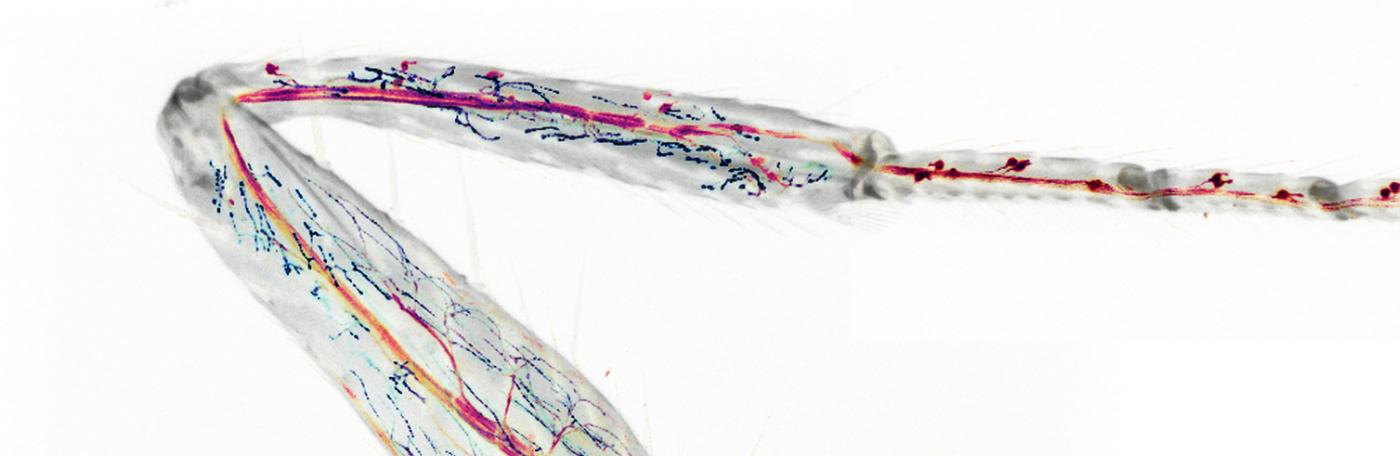
We are studying how the neural circuitry that is required for coordinated walking in adult flies is constructed during development, and how it functions in the adult.
News
New lab members: Ryan Kittle and Kristi Xing
The lab welcomes Ryan Kittle, from the Genetics and Development Department. Ryan is doing his first rotation in the lab this fall.
We also welcome Kristi Xing, a Barnard undergrad, who will be working on fly neuroscience!
Farewell to Siqian
This month we say farewell and best of luck to Siqian Feng, who will be starting his own research group at the School of Life Science and Technology of Southeast University, Nanjing, China. We will greatly miss Siqian's creativity, his consistent goal to achieve excellence, and his ample willingness to give his time and expert advice to anyone who asks. But we also look forward to continued interactions with the Feng lab in the future!
Ross Munce is awarded an F31 grant from the NIH
Congratulations to Ross on getting this award!!!
Photos

Recent Lab dinner, saying farewell to Siqian!
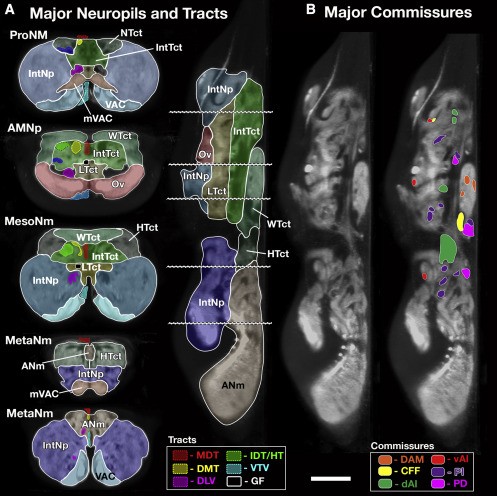
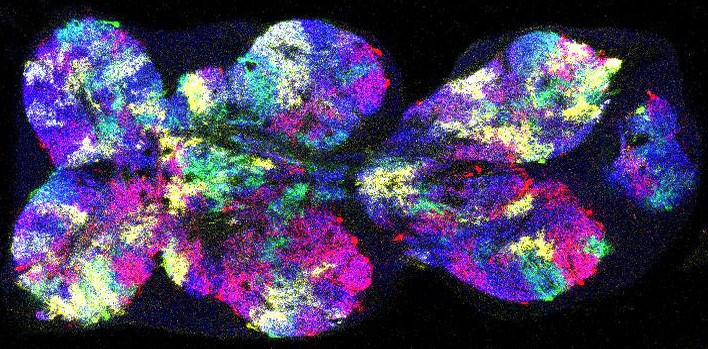
Neuropils of the Drosophila VNC
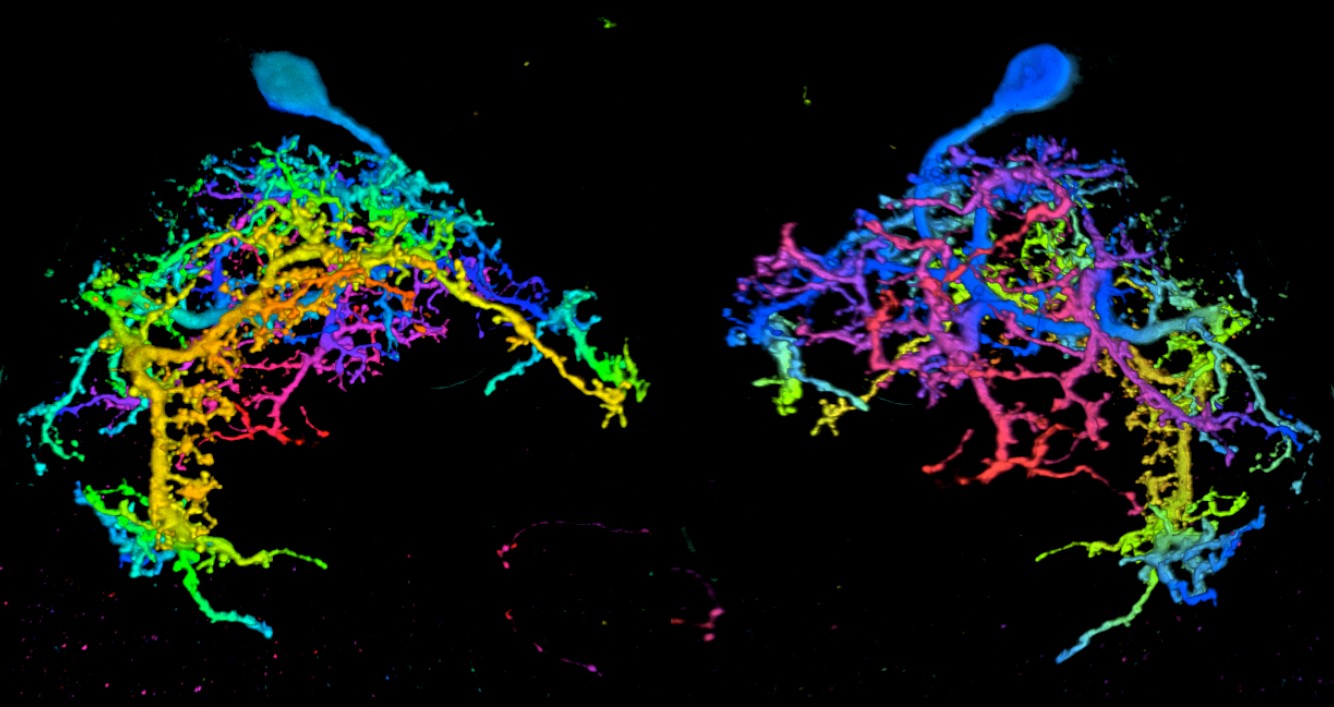
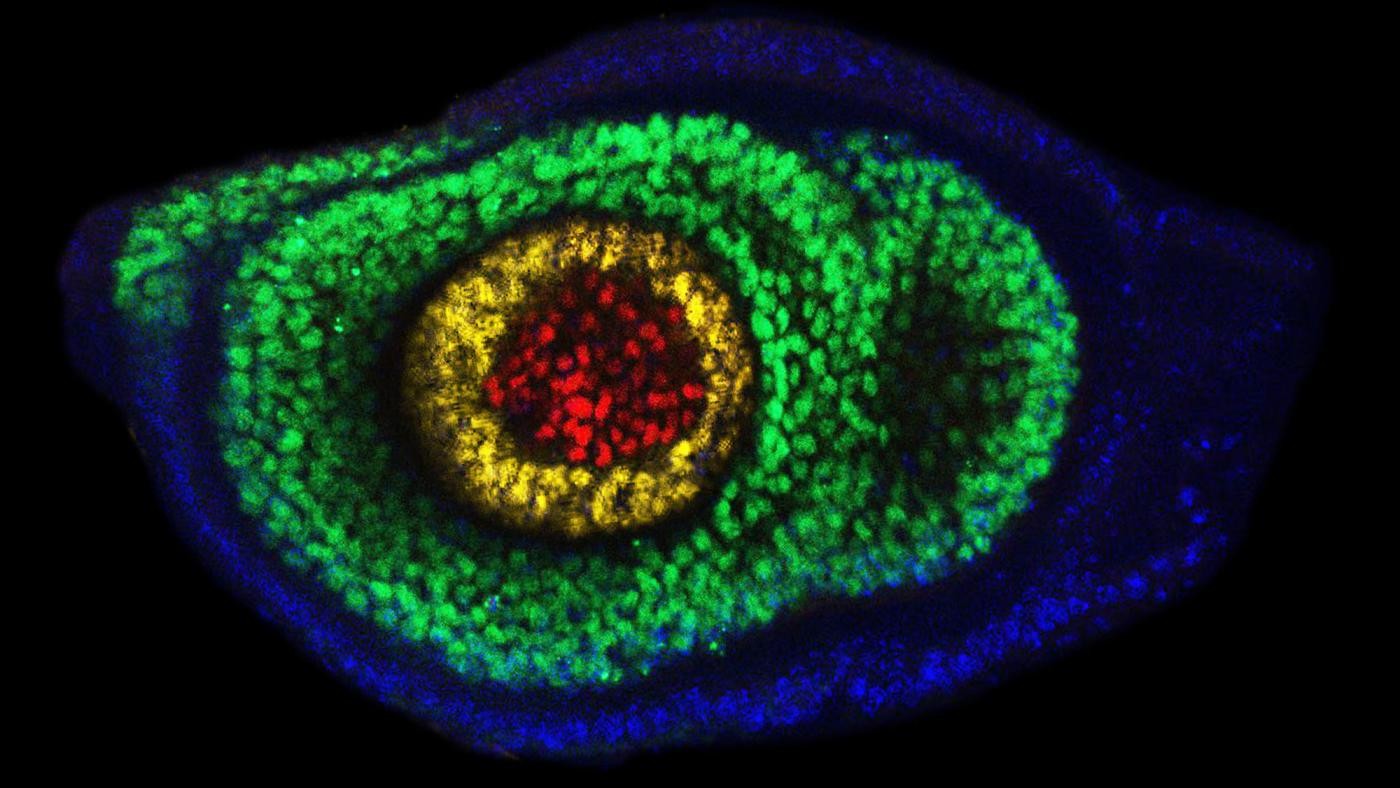
Dendrites and cell bodies of a pair of tarsal depressor motor neurons