2025
FETCH enables fluorescent labeling of membrane proteins in vivo with spatiotemporal control in Drosophila
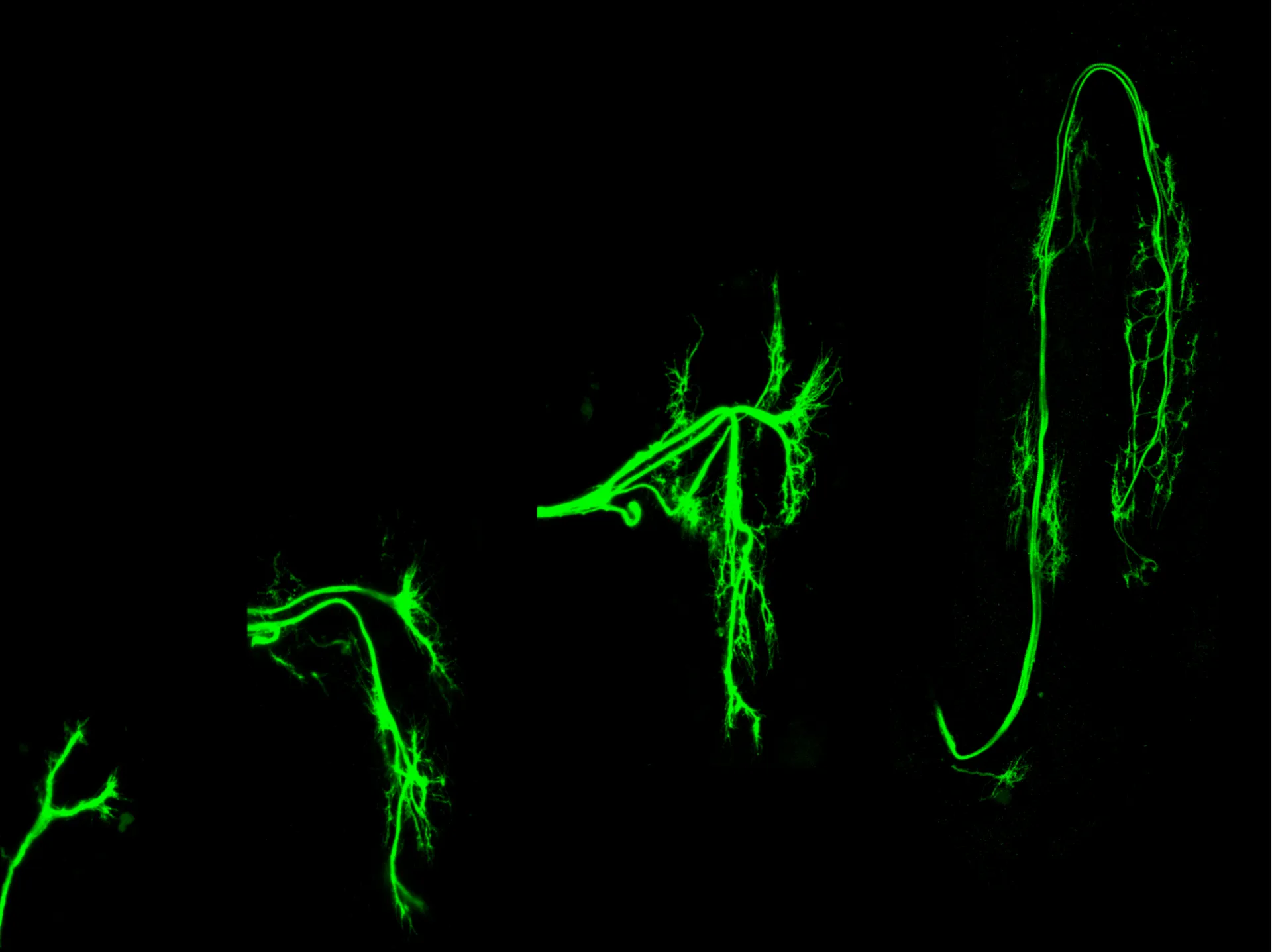
A critical affinity window for IgSF proteins DIP-α and Dpr10 is required for proper motor neuron arborization
Predicting the DNA binding specificity of transcription factor mutants using family-level biophysically interpretable machine learning
Decoding neuronal wiring by joint inference of cell identity and synaptic connectivity
Members of the DIP and Dpr adhesion protein families use cis inhibition to shape neural development in Drosophila
2024

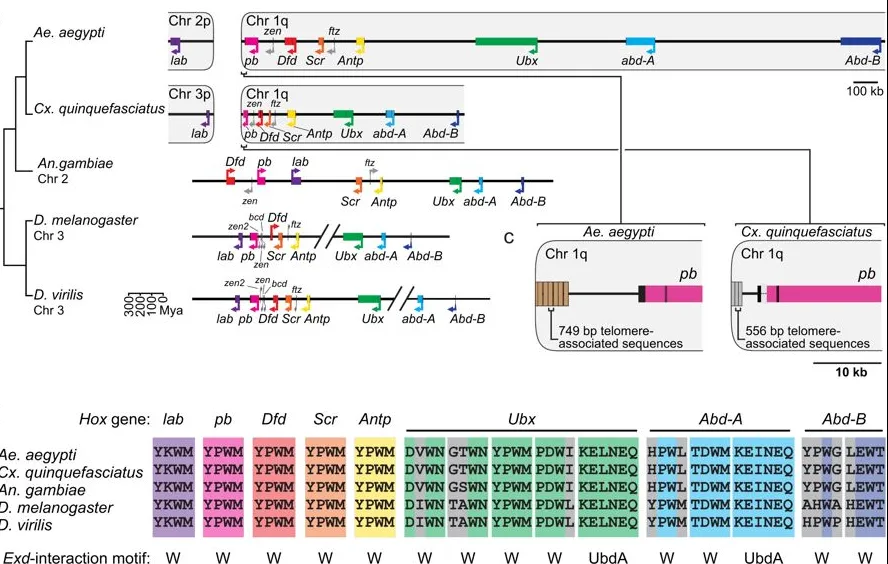
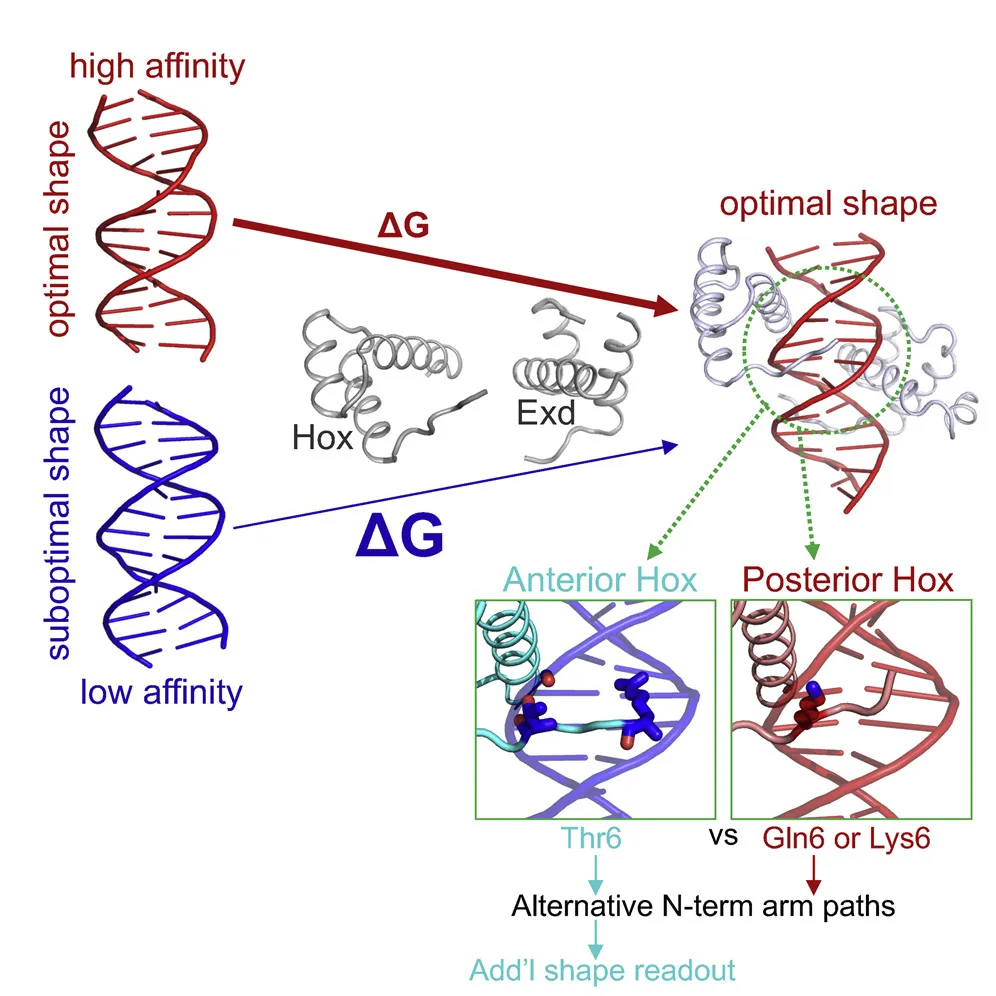
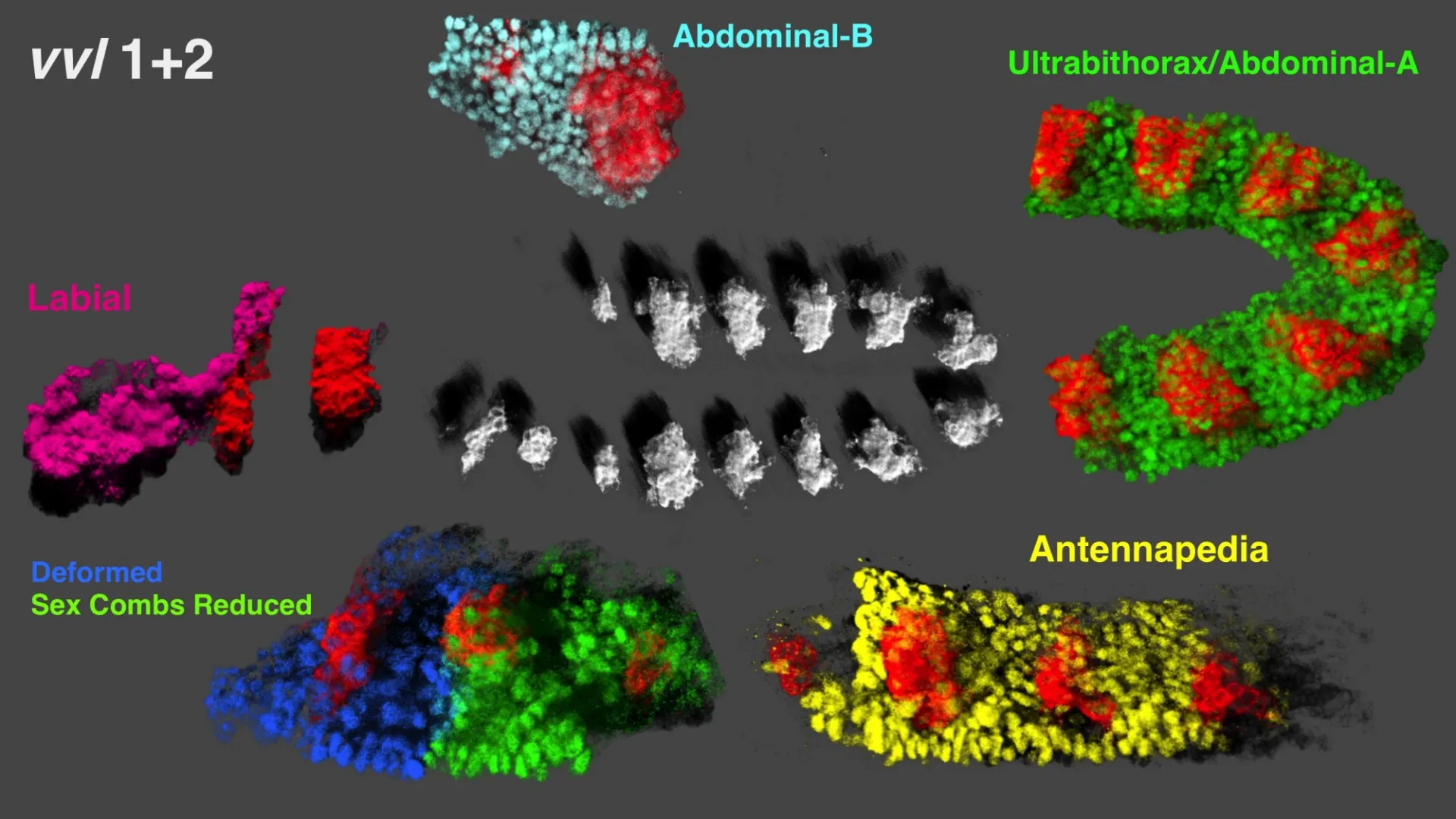
HOX GENES: THE ORIGINAL BODY BUILDERS
2022
Transcription factor paralogs orchestrate alternative gene regulatory networks by context dependent cooperation with multiple cofactors
SpyChIP identifies cell type-specific transcription factor occupancy from complex tissues
2021

Cell-type-specific Hox regulatory strategies orchestrate tissue identity
Homothorax controls a binary Rhodopsin switch in Drosophila ocelli
Scarless engineering of the Drosophila genome near any site-specific integration site.
2020
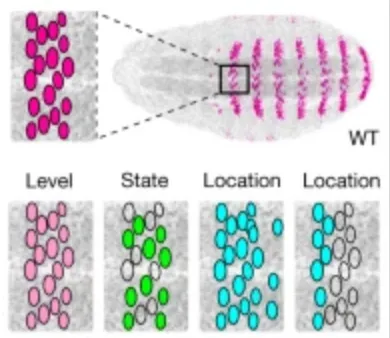
Dense and pleiotropic regulatory information in a developmental enhancer
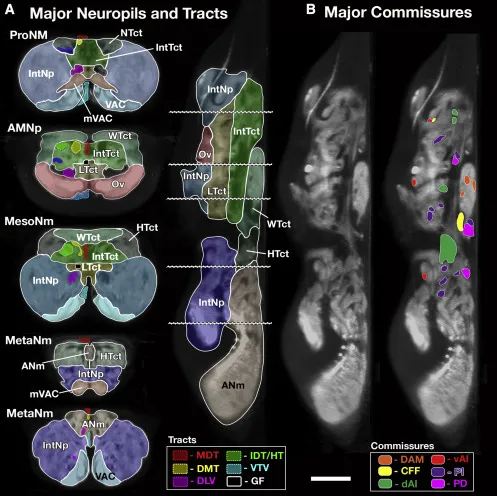
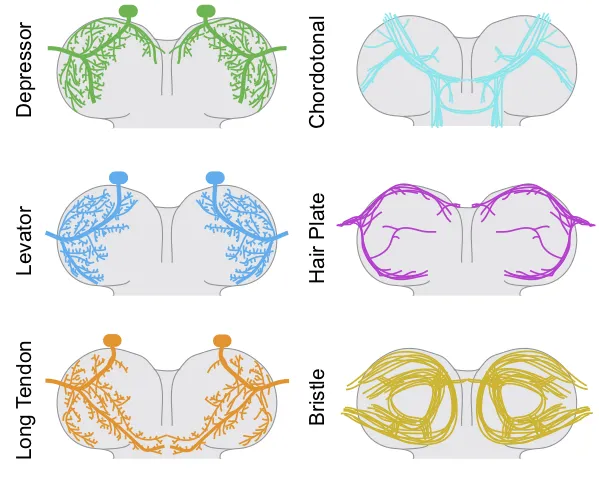
A size principle for recruitment of Drosophila leg motor neurons
Control of tissue morphogenesis by the HOX gene Ultrabithorax
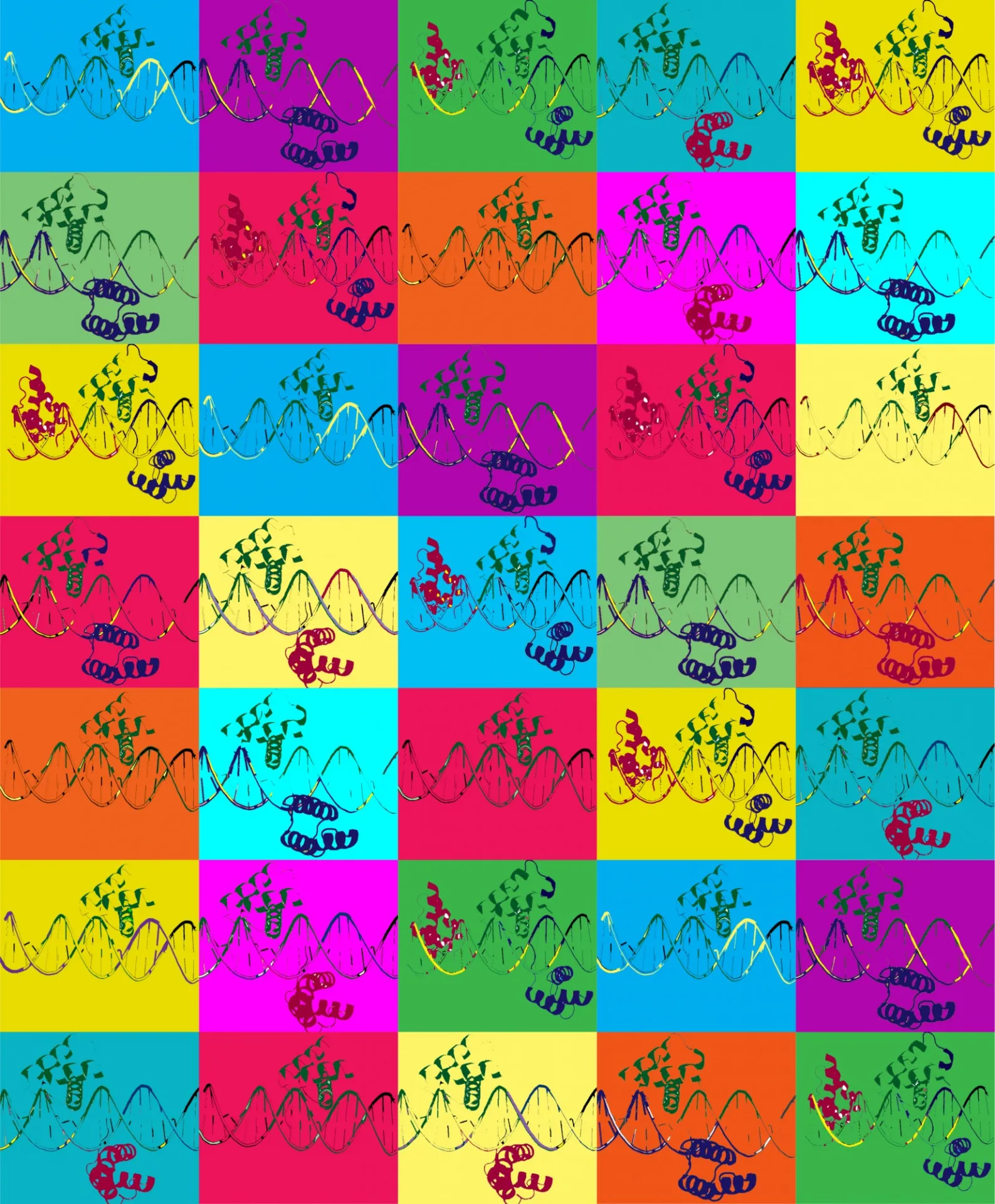

2019
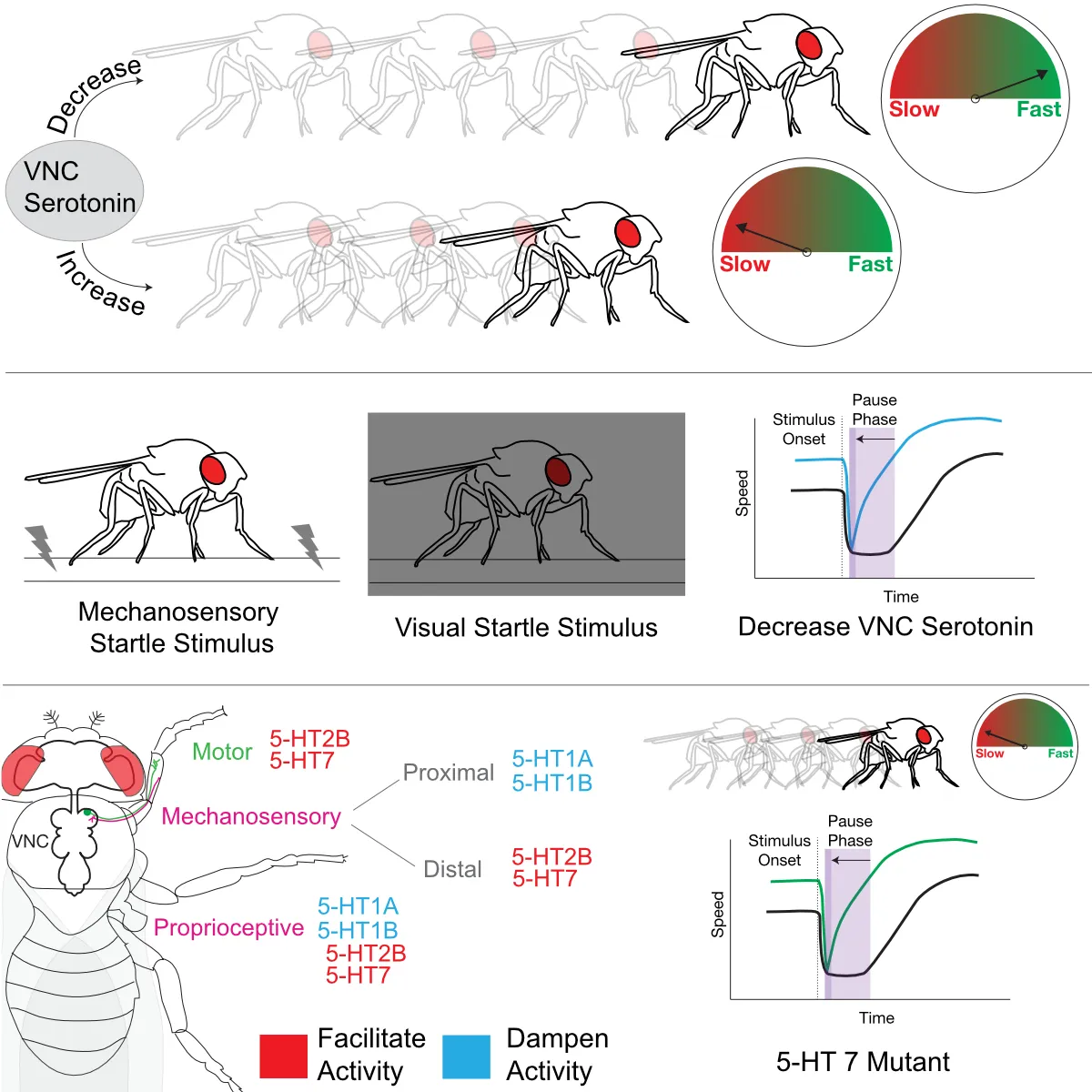
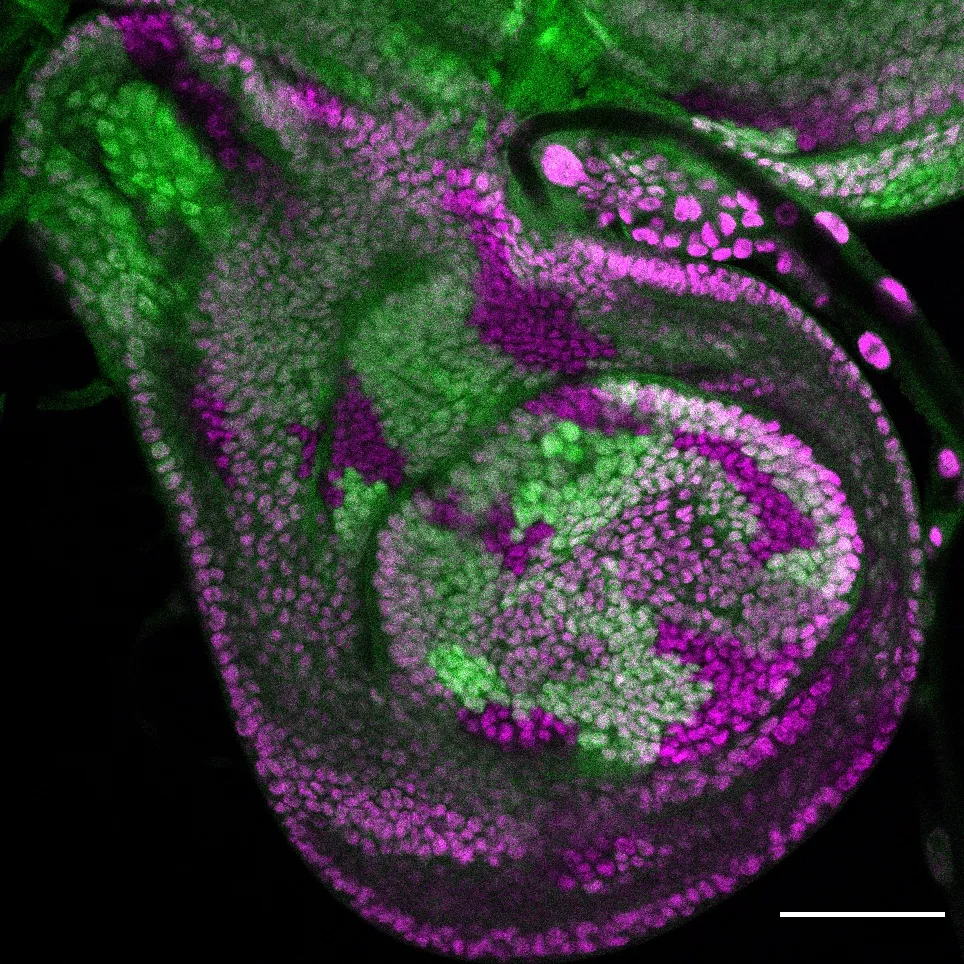
A role for low affinity binding sites in fine tuning the levels of transcription
Low-Affinity Binding Sites and the Transcription Factor Specificity Paradox in Eukaryotes
No results
There are no publications with the provided filters.